Ordering
*link will take you to our exclusive distribution partner site
*link will take you to our exclusive distribution partner site
The SensiFAST Probe No-ROX One-Step Kit has been optimized for fast, efficient, unbiased cDNA synthesis and subsequent highly-sensitive, reproducible real-time PCR detection in a single tube. SensiFAST Probe One-Step has been optimized to deliver excellent results in both singleplex and multiplex assays.
An antibody-mediated hot-start DNA polymerase promotes rapid activation and supports highly-specific amplification, which in turn improves assay sensitivity and dynamic range. A combination of the latest advances in buffer chemistry and PCR enhancers confer superior assay performance under fast thermal cycling conditions. The inclusion of separate RiboSafe Inhibitor ensures accuracy by protecting RNA targets from RNase degradation.
The SensiFAST™ Probe No-ROX One-Step Kit has been validated on all commonly-used real-time instruments that do not require the passive reference dye ROX. SensiFAST Probe One-Step has been formulated for use with dual-labelled probes, including TaqMan®, Scorpions® and molecular beacon probes.
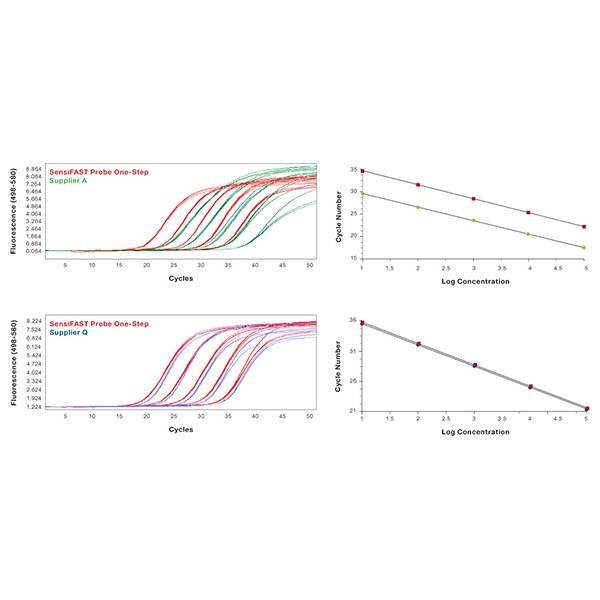
Mouse β-actin was amplified in triplicate using gene-specific primers and TaqMan probe according to each manufacturer's protocol, from 10-fold serial dilution of RNA, using either SensiFAST Probe One-Step Kit (red) or similar kits from Suppliers A (green) and Q (blue). The SensiFAST Probe One-Step Kit gives the lowest Ct values while maintaining reproducibility of technical replicates.
A fragment of the mouse β-actin gene was amplified in triplicate from a 10-fold serial dilution of mouse 3T3 total RNA using SensiFAST Probe One-Step Kit, gene-specific primers and a TaqMan probe, under fast thermal cycling conditions. The amplification plot shows excellent assay reproducibility.
A 100-fold dilution of human cDNA was targeted with 4 different gene-specific assays, in either singleplex reactions (dark coloured line) or quadruplex assays (light coloured line), using the SensiFAST Probe One-Step Kit and similar kits from Suppliers Q and T. The results demonstrate that SensiFAST Probe One-Step Kit is very efficient in multiplexing, as reactions containing single and four probes give the same Ct value, unlike the kits from Suppliers Q and T.
|
Reagent |
100 x 20 µL Reactions |
500 x 20 µL Reactions |
|
SensiFAST Probe No-ROX One-Step Mix (2x) |
1 x 1 mL |
5 x 1 mL |
|
RiboSafe RNase Inhibitor |
1 x 40 µL |
1 x 200 µL |
|
Reverse transcriptase |
1 x 20 µL |
1 x 100 µL |
|
DEPC-H2O |
1 x 1.8 mL |
2 x 1.8 mL |
All kit components should be stored at -20°C upon receipt for optimum stability. Repeated freeze/thaw cycles should be avoided.
When stored under the recommended conditions and handled correctly, full activity of the kit is retained until the expiry date indicated on the outer box label.
Shipped on dry/blue ice.