Ordering
*link will take you to our exclusive distribution partner site
*link will take you to our exclusive distribution partner site
MyFi™ has been developed to give reliable amplification of targets up to 10 kb from challenging and complex targets. MyFi shows improved tolerance to PCR inhibitors, thereby enabling reliable detection from samples from which DNA is difficult to purify. Furthermore a unique buffer system and enzyme blend promote highly sensitive amplification, ideal for low-copy number targets. The inclusion of MyTaq HS means MyFi generates PCR products with 3’-A overhangs making it suitable for TA cloning. MyFi has the added convenience of room temperature reaction assembly, to avoid non-specific amplification and primer-dimer formation.
An advanced buffer system and enzyme blend combine to give increased target affinity, ideal for amplification of cDNA libraries, complex genomic fragments and GC-rich targets.
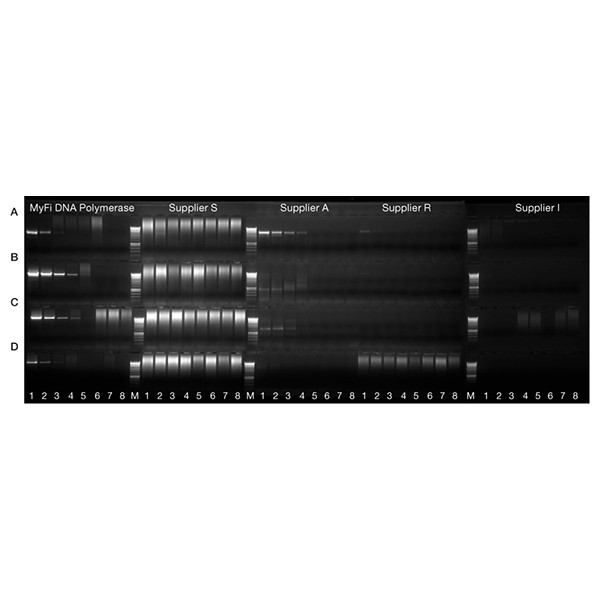
Greater reliability with GC-rich DNA
A 525 bp fragment of the 62% GC-rich human epidermal growth factor receptor (EGFR) gene (A), a 750 bp fragment of the 64% GC-rich translation factor p64 (myc) gene (B), a 900 bp fragment of 43% GC-rich angiotensin II receptor type I (AGTR1) gene (C), a 1.2 kb fragment of 62% GC-rich EGFR gene (D) were amplified using MyFi DNA Polymerase and similar polymerases from other suppliers. A five-fold serial dilution of human genomic DNA (25 ng - 8 pg, 1.6 pg and 0 pg, lanes 1-8 respectively) was used as template. Marker is HyperLadder 1kb (M). The results demonstrate that that MyFi delivers more reliable amplification of challenging targets.
Amplification of complex DNA up to 10kb
A 3.9 kb fragment of a-1-antitrypsin (AT-R3) gene (A), and a 7.0 kb (B), 9.0 kb (C) and 10.0 kb (D) fragment of human (ß-globin) HbG gene, were amplified using MyFi DNA Polymerase and similar DNA Polymerases from other suppliers. A serial dilution of human genomic DNA (5 ng, 1 ng, 200 pg, 40 pg, 8 pg, 1.6 pg, 0.32 pg and 0 pg, Lanes 1-8 respectively) was used as template. Marker is HyperLadder 1kb (M). The results demonstrate that MyFi can be used to amplify targets up to 10 kb from even very challenging templates.
|
Reagent |
250 Units |
500 Units |
2500 Units |
|
MyFi DNA Polymerase |
1 x 125 µL |
1 x 250 µL |
2 x 625 µL |
|
5x MyFi Reaction Buffer |
1 x 625 µL |
1 x 1.25 mL |
5 x 1.25 mL |
BIO-21117: 125 x 25 µL Reactions: 1 x 125 µL
BIO-21118: 250 x 25 µL Reactions: 1 x 250 µL
BIO-21119: 1250 x 25 µL Reactions: 2 x 625 µL
2 u/µL
All components should be stored at -20°C upon receipt for optimum stability. Repeated freeze/thaw cycles should be avoided.
When stored under the recommended conditions and handled correctly, full activity of the reagents is retained until the expiry date indicated on the outer box label.
On Dry Ice or Blue Ice.
| Observation | Recommended Solution(s) |
| No or low PCR yield | Enzyme concentration too low – increase the amount of enzyme in 0.5 U increments. |
| Primers degraded – check quality and age of the primers. | |
| Magnesium concentration too low – increase concentration in 0.25 mM increments with a starting concentration of 1.75 mM. | |
| Primer concentration not optimized. Titrate primer concentration (0.3-1 µM); ensuring that both primers have the same concentration. | |
| Template concentration too low – Increase concentration of template. | |
| Perform a positive control to ensure that the enzyme, dNTPs and buffers are not degraded and/or contaminated. | |
| Multiple Bands | Primer annealing temperature too low. Increase annealing temperature. Primer annealing should be at least 5°C below the calculated Tm of primers. |
| Prepare master mixes on ice or use a heat-activated polymerase. | |
| For problems with low specificity. Try adding 3% DMSO (not supplied) to improve specificity. | |
| Smearing or artifacts | Template concentration too high. Prepare serial dilutions of template. |
| Too many cycles. Reduce the cycle number by 3-5 to remove non-specific bands. | |
| Enzyme concentration too high - decrease the amount of enzyme in 0.5 U increments. | |
| Extension time too long. Reduce extension time in 0.5-1 minute increments. |
Please click here in order to request your sample. You will receive an email confirmation within two business days with delivery details.